|
|
Developmental Biology - Cell Structure
The Fireworks of life
Biologists reverse engineer to find how microtubules hold up cell walls and make spindles...
Imagine standing in a lumberyard and being told to build a house - without blueprints or instructions of any kind. The materials are there, but you haven't the first idea how to get from point A to point B.
That was the situation for Princeton biologists wanting to build the microtubule skeleton of a cell.
According to Sabine Petry, assistant professor of molecular biology: "We didn't think it was possible." For years, Petry and her lab dazzled the biological world with videos of "the fireworks of life." But, getting the recipe of how fireworks are made? "We'd imagined and brainstormed about it for five years," said Petry. Showing microtubules as they branch and grow microscopic structures, Petry's team eventually, painstakingly determined each fireworks' component, one protein at a time.
However, journal reviewers specified their research couldn't be published unless proven experimentally. It would later appear in the May 8, issue of eLife.
Building a house without blueprints
While Petry's team had identified the components necessary to build microtubules, they hadn't put together the recipe for how to assemble those pieces into the fireworks of growing spindles, branching at the molecular level.
How did it happen, exactly? was the question nagging at Akanksha Thawani PhD, chemical and biological engineering graduate student [who recently won the prestigious Charlotte Elizabeth Proctor Fellowship for graduate students in their final year]. "We start from no microtubules at all, and then, within 15 minutes, we have these beautiful structures. How do you generate a structure from those nanometer-sized proteins? What was it about, their binding kinetics, or the organization that would result in the structures that we see?"
Thawani is uniquely positioned to tackle these questions, as a chemical engineering/physics student, and a student of molecular biology. Thawani essentially invented a new subspecialty in-between the three fields. "At the intersections between disciplines - that is where the next, best science is," she explains.
Each of three principal investigators (PIs) have research labs in three unrelated fields. Petry in biology; Stone in engineering; and Joshua Shaevitz, a professor of physics and the Lewis-Sigler Institute for Integrative Genomics. Thawani realized the need to optically isolate branching microtubules from a sea of background molecules using total internal reflection fluorescence (TIRF), a technique the Petry lab had developed.
Every pixel recorded by the camera included thousands of molecules. Thawani had to disaggregate this visual data into single-molecule observations, requiring months of complicated image analysis with help from Joshua Shaevitz who has spent years on image analysis.
Ultimately, Shaevitz measured exactly when and where a single protein binds to an existing microtubule to begin a new branch. The model also notes the rate of that branch's growth, by following one molecule at a time.
"By looking at individual molecules, we can literally watch the assembly piece by piece as it happens."
Joshua Shaevitz PhD, Professor of Physics, Lewis-Sigler Institute for Integrative Genomics, and Co-Director, National Science Foundation Center for Physics of Biological Function.
Thawani then created a computer model following Shaevitz' parameters. She then tested various sequences that the researchers had brainstormed over for years, and ruled out all but one model.
The research team now had the ingredients - proteins called TPX2, augmin and y-TuRC, as well as the sequence of steps. But, the computer couldn't tell them in what order to add proteins - or when.
The Final Twist
Experiments required for publication revealed another problem. Thawani and Petry's expectations were backwards. "We went in thinking augmin had to be first and then TPX2. It turned out to be the other way around," Thawani said. "That was the twist." With this discovery, they had their complete recipe to generate microtubule fireworks.
If TPX2 is deposited on existing microtubules, followed by binding augmin with y-TuRC, then new microtubules will nucleate and branch.
As a final step, researchers confirmed that proteins bind with precisely the speed predicted by Thawani's computer model. "That was the third breakthrough," Petry says, "that the numbers matched as was predicted by her computer model, was also true for the biology."
"This work from Petry is really an important addition that will help drive the field forward. I think that this work, in combination with results from my group and from Jan Brugués (at the Max Planck Institute of Molecular Cell Biology and Genetics in Dresden), have really clarified the 'rules' for microtubule nucleation in spindles. The next step will be to figure out the molecular processes that govern those rules. Petry and colleges have setup a system that should really help to do that."
Daniel Needleman PhD, the Gordon McKay Professor of Applied Physics, Professor of Molecular and Cellular Biology, Harvard University, Massachusetts, USA.
Abstract
Sequencing-based studies have identified novel risk genes associated with severe epilepsies and revealed an excess of rare deleterious variation in less-severe forms of epilepsy. To identify the shared and distinct ultra-rare genetic risk factors for different types of epilepsies, we performed a whole-exome sequencing (WES) analysis of 9,170 epilepsy-affected individuals and 8,436 controls of European ancestry. We focused on three phenotypic groups: severe developmental and epileptic encephalopathies (DEEs), genetic generalized epilepsy (GGE), and non-acquired focal epilepsy (NAFE). We observed that compared to controls, individuals with any type of epilepsy carried an excess of ultra-rare, deleterious variants in constrained genes and in genes previously associated with epilepsy; we saw the strongest enrichment in individuals with DEEs and the least strong in individuals with NAFE. Moreover, we found that inhibitory GABAA receptor genes were enriched for missense variants across all three classes of epilepsy, whereas no enrichment was seen in excitatory receptor genes. The larger gene groups for the GABAergic pathway or cation channels also showed a significant mutational burden in DEEs and GGE. Although no single gene surpassed exome-wide significance among individuals with GGE or NAFE, highly constrained genes and genes encoding ion channels were among the lead associations; such genes included CACNA1G, EEF1A2, and GABRG2 for GGE and LGI1, TRIM3, and GABRG2 for NAFE. Our study, the largest epilepsy WES study to date, confirms a convergence in the genetics of severe and less-severe epilepsies associated with ultra-rare coding variation, and it highlights a ubiquitous role for GABAergic inhibition in epilepsy etiology.
Authors
Akanksha Thawani, Howard A. Stone, Joshua W. Shaevitz and Sabine Petry.
Acknowledgements
This work was supported by the American Heart Association predoctoral fellowship 17PRE33660328 (to AT), the NIH New Innovator Award 1DP2GM123493, Pew Scholars Program in the Biomedical Sciences 00027340, David and Lucile Packard Foundation 2014-40376 (all to SP), and the Center for the Physics of Biological Function sponsored by the National Science Foundation grant PHY-1734030.
Return to top of page.
| |
|
Aug 20 2019 Fetal Timeline Maternal Timeline News
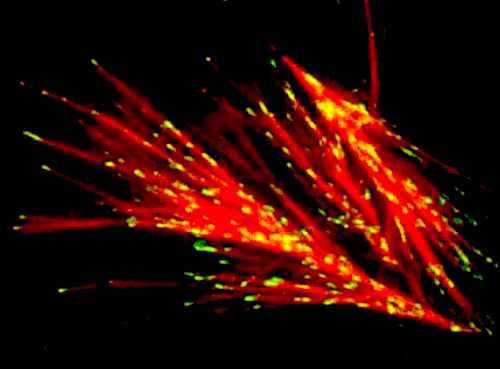 A scene from the growth and branching of microtubules video 'The Fireworks of Life' shows how microscopic structures grow in the cell. Scientists reverse engineered the recipe for building these fireworks, capturing their growth in seconds on the scale bar at 10 µm (a human hair is 50 to 100 µm across). CREDIT Akanksha Thawani, Princeton University
|



