|
|
Developmental Biology - Neuropsychiatric Diseases
RNA Creates Fetal Map of Neuropsychiatric Disorders
Studying age-specific genetic effects may open more doors on the mechanisms behind brain disorders...
From early prenatal development through childhood, the prefrontal cortex of the human brain undergoes an avalanche of activity. According to a new genetic analysis led by researchers at Yale University and the University of California-San Francisco (UCSF), the prenatal period may also contain seeds of neuropsychiatric illness such as autism spectrum disorder and schizophrenia.
Previous studies have identified DNA variations linked to neuropsychiatric illnesses. But, only when those variations triggered change in the dorsal lateral prefrontal cortex linked to neuropsychiatric, cognitive, and emotional disorders. The new study, published April 7 in the journal Cell Reports, added a new dimension to prior research.
Scientists measuring the amount of RNA were able to provide a picture of overall gene activity. In 176 tissue samples taken across a variety of developmental stages, they determined how and when DNA variations influence brain function.
"This is the first large cohort to profile DNA and RNA both in prenatal and postnatal human brain samples, making it an unprecedented resource for understanding how individual genetic differences might lead to functional differences."
Sirisha Pochareddy PhD, Associate Research Scientist, Department of Neuroscience and Kavli Institute for Neuroscience, Yale School of Medicine, New Haven, Connecticut, USA and co-lead author of the study.
Researchers tracked thousands of variants associated with thousands of genes across the entire genome.
This level of sophistication allowed scientists to identify groups of genes regulating distinct biological processes that can lead to disease.
Understanding how genetic variation and changes in function are linked points out alterations in brain development and can pick out schizophrenia and autism later in life, say the authors.
"Human brain development is an incredibly complex and dynamic process, and any disruption along the way can have profound consequences on later brain function.
Interestingly, we found some genetic variants have stronger effects on RNA expression before birth other variants have strongest effects after birth."
Donna Werling PhD, University of Wisconsin-Madison, formerly with UCSF.
Abstract Highlights
Whole-genome and RNA sequencing across human
prefrontal cortex development
Gene-specific developmental trajectories characterize the late-fetal transition
Identification of constant, prenatal-predominant, and postnatal-predominant eQTLs
Integrated analysis implicates genes in loci associated with educational attainment
Authors
Donna M. Werling, Sirisha Pochareddy, Jinmyung Choi, Joon-Yong An, Brooke Sheppard, Minshi Peng, Zhen Li, Claudia Dastmalchi, Gabriel Santpere, Andre´ M.M. Sousa, Andrew T.N. Tebbenkamp, Navjot Kaur, Forrest O. Gulden, Michael S. Breen, Lindsay Liang, Michael C. Gilson, Xuefang Zhao, Shan Dong, Lambertus Klei, A. Ercument Cicek, Joseph D. Buxbaum, Homa Adle-Biassette.
Acknowledgements
Data were generated as part of the PsychENCODE Consortium, supported
by U01MH103339, U01MH103365, U01MH103392, U01MH103340,
U01MH103346, R01MH105472, R01MH094714, R01MH105898,
R21MH102791, R21MH105881, R21MH103877, and P50MH106934 awarded
to Schahram Akbarian (Icahn School of Medicine at Mount Sinai), Gregory
Crawford (Duke), Stella Dracheva (Icahn School of Medicine at Mount Sinai),
Peggy Farnham (USC), Mark Gerstein (Yale), Daniel Geschwind (UCLA),
Thomas M. Hyde (LIBD), Andrew Jaffe (LIBD), James A. Knowles (USC), Chunyu
Liu (UIC), Dalila Pinto (Icahn School of Medicine at Mount Sinai), Nenad Sestan
(Yale), Pamela Sklar (Icahn School of Medicine at Mount Sinai), Matthew
State (UCSF), Patrick Sullivan (UNC), Flora Vaccarino (Yale), Sherman Weissman
(Yale), Kevin White (UChicago), and Peter Zandi (JHU). This work was
supported by funding provided by Autism Science Foundation postdoctoral
fellowships (to D.W. and J.-Y.A.) and a research award (to S.J.S.); Simons
Foundation Autism Research Initiative (SFARI) grants 574598 (to S.J.S.),
402281 (to S.J.S., M.W.S., B.D., and K.R.), and 573206 (to M.E.T.); National
Institute for Mental Health (NIMH) grants R01 MH109901 (to S.J.S. and
M.W.S.), R01 MH110928 (to S.J.S. and M.W.S.), U01 MH103339
(to M.W.S.), R01 MH111662 (to S.J.S. and M.W.S.), U01 MH105575 (to
M.W.S.), U01 MH106874 (to N.S.), P50 MH106934 (to N.S.), R01 MH109904
(to N.S.), R01 MH110926 (to N.S.), U01 MH116488 (to N.S.), R37 MH057881
(to B.D.), and R01 MH115957 (to M.E.T.); National Institute of Child Health
and Human Development (NICHD) grants R24 HD000836 (to I.A.G.), R01
HD081256 (to M.E.T.), and R01 HD096326 (to M.E.T.); National Heart, Lung,
and Blood Institute (NHLBI) grant K25HL121295 (to N.A.Z.); National Human
Genome Research Institute (NHGRI) grants U01HG009080 (to N.A.Z.) and
R01HG006399 (to N.A.Z.); National Cancer Institute (NCI) grant
R01CA227237 (to N.A.Z.); National Institute of Dental and Craniofacial
Research (NIDCR) grant R03DE025665 (to N.A.Z.); United States Department
of Defense (DoD) grant W81XWH-16-2-0018 (to N.A.Z.); and National
Research Foundation of Korea grant 2019M3E5D3073568 (to J.-Y.A.). The
project that gave rise to these results also received the support of a fellowship
from la Caixa Foundation (ID 100010434 to G.S.). The fellowship code is
LCF/BQ/PI19/11690010. We thank Thomas Lehner, Anjene´ Addington, Geetha
Senthil, and Alexander Arguello at the NIMH for supporting the PsychENCODE
Consortium, the Yale Center for Genome Analysis for generating the
RNA-seq data, GENEWIZ for generating the WGS data, and Sentieon for
use of a computationally efficient implementation of GATK Haplotype Caller.
Return to top of page.
| |
|
Apr 13 2020 Fetal Timeline Maternal Timeline News
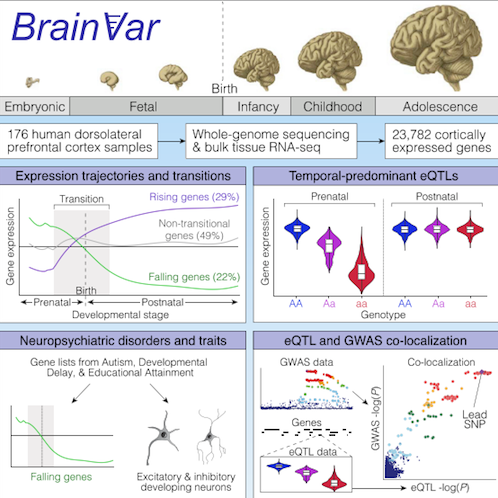 After analyzing gene expression across human cerebral cortical development, scientists identified specific genes can be associated with neuropsychiatric traits and disorders.
|



