|
|
The silence of the genes
Within each of our cells, we have two copies of each gene we inherited, one from our mother and one from our father. If one of those two genes is 'turned off', or silenced, it can affect health.
Research led by Keiji Tanimoto DDS, PhD in Cancer Research and Molecular Biology at the University of Tsukuba, Japan, has brought a closer understanding of the underlying phenomenon of genomic imprinting. In this intriguing event, one copy of a gene is 'turned off'— or silenced — depending on whether it was derived from the mother or the father.
The research identified a segment of DNA essential in imprinting for closely linked Igf2/H19 genes, two of the first imprinted genes to be discovered. If these genes are incorrectly imprinted, the results can be overgrowth (Beckwith-Wiedemann) or dwarfism (Silver Russell) syndromes. The genes also have a role in some kidney and liver cancers.
Of those two inherited copies of every gene - one from mother, one from father - most are active and working. In a small number of genes, however, one is turned off depending on its parent of origin. These genes are called imprinted because one copy is tagged via modifications to its DNA — either egg or sperm. The mechanisms for imprinting are not fully understood, but involve modifications to DNA being removed and reset during creation of the egg or sperm. These modifications are ' epigenetic' (or outside the gene), as the DNA sequence itself is not altered.
Normally we only have one copy of a silenced gene.
If an error occurs in the imprinting process, we might end up with two active copies of a gene or two inactive or silenced copies.
This could lead to abnormal development, diseases, cancer and various behavioral disorders, such as the neurodevelopmental disorders of Prader-Willi and Angelman syndromes.
Imprinting disorders have also been linked to diabetes.
Reported in the latest volume of the journal Development, the scientists investigated when and how imprinted Igf2/H19 genes in mice are modified in early development to further understand the mechanisms underlying the imprinting process.
The Igf2/H19 genes in mice are two of the best understood imprinted genes with the same imprinting pattern being found in human equivalents, making the mice ideal candidates for further research.
Imprinted genes are controlled by nearby DNA sequences, called Imprinting Control Regions (ICRs). ICRs have parental-specific modifications, including DNA methylation — a chemical reaction that attaches small molecules called methyl groups to certain segments of the DNA. In genes undergoing genomic imprinting, methylation is one way a gene's parent of origin is marked during the formation of egg and sperm.
ICRs are tagged by proteins called DNA methyltransferases (DMTs) during egg and sperm development. Methylation is faithfully maintained on the imprinted gene throughout development, even in early embryos where genomes undergo extensive reprogramming, including removal of methylation from the DNA, like a scar.
The H19 ICR in the mouse is DNA-methylated by two DMTs, Dnmt3a and Dnmt3L, during sperm formation. This methylation is maintained on the paternally-derived copy of the Igf2/H19 genes following fertilization, rendering these genes 'imprinted'. The research team were interested to discover when and how this methylation is so faithfully maintained.
Using an elegant genetic strategy involving cross-breeding several strains of transgenic mice, they found paternal-specific methylation of H19 ICR begins soon after fertilization, but also that maternally-supplied DNA methyltransferases Dnmt3a and Dnmt3L are required. This is the first example to demonstrate a role for Dnmt3L in early embryogenesis.
Even when the methylation of the mouse H19 ICR was only partially obstructed in sperm, paternal H19 ICR exhibited methylation. It therefore seems likely H19 ICR acquired an unknown epigenetic mark or tag, not DNA methylation, during sperm formation and after fertilization the de novo DNA methylation machinery recognized this mark.
Possible candidates for the epigenetic marks acquired during sperm formation are modifications or variants in histones, the proteins around which DNA is wrapped.
The team was able to narrow down the sequence for postfertilization acquisition of methylation to a sequence located at the beginning of the ICR, the so-called 5' region. Deletion of this 5' region results in a partial loss of methylation on the paternal gene, abnormal expression of H19 and its neighboring imprinted gene Igf2, and retards growth.
This is the first example of a regulatory DNA sequence able to regulate methylation of paternal H19 ICR, all other regulatory sequences, so far, maintain the unmethylated state of the maternal H19 ICR after implantation.
"These results demonstrate that this segment of the H19 ICR is essential for de novo post-fertilization DNA methylation, and that this activity contributes to the maintenance of imprinted methylation on H19 ICR during early embryogenesis."
Keiji Tanimoto DDS, PhD, Cancer Research and Molecular Biology, Hiroshima University, Hiroshima, Japan
One of the next steps is to find the primary mark that instructs imprinted DNA methylation in early embryos in genes responsible for imprinting disorders in humans.
This information may have therapeutic benefits by allowing us, for example, to induce re-expression of a silent parental gene copy in patients in order to improve their symptoms.
Abstract
Genomic imprinting is a major monoallelic gene expression regulatory mechanism in mammals, and depends on gamete-specific DNA methylation of specialized cis-regulatory elements called imprinting control regions (ICRs). Allele-specific DNA methylation of the ICRs is faithfully maintained at the imprinted loci throughout development, even in early embryos where genomes undergo extensive epigenetic reprogramming, including DNA demethylation, to acquire totipotency. We previously found that an ectopically introduced H19 ICR fragment in transgenic mice acquired paternal allele-specific methylation in the somatic cells of offspring, while it was not methylated in sperm, suggesting its gametic and postfertilization modifications are separable events. We hypothesized that this latter activity might contribute to maintenance of the methylation imprint in early embryos. Here we demonstrate that methylation of the paternally inherited transgenic H19 ICR commences soon after fertilization in a maternal Dnmt3a- and Dnmt3L-dependent manner. When its germline methylation was partially obstructed by insertion of insulator sequences, the endogenous, paternal H19 ICR also exhibited postfertilization methylation. Finally, we refined the responsible sequences for this activity in transgenic mice, and found that deletion of the 5' segment of the endogenous paternal H19 ICR decreased its methylation after fertilization, attenuated Igf2 gene expression. These results demonstrate that this segment of the H19 ICR is essential for its de novo post-fertilization DNA methylation, and that this activity contributes to the maintenance of imprinted methylation at the endogenous H19 ICR during early embryogenesis.
Universities involved in this study are: the University of Tsukuba, the Nagahama Institute of Bio-Science and Technology, Osaka University and the National Research Institute for Child Health and Development.
Received May 2, 2015.
Accepted September 15, 2015.
Return to top of page
|
|
|
Dec 3, 2015 Fetal Timeline Maternal Timeline News News Archive
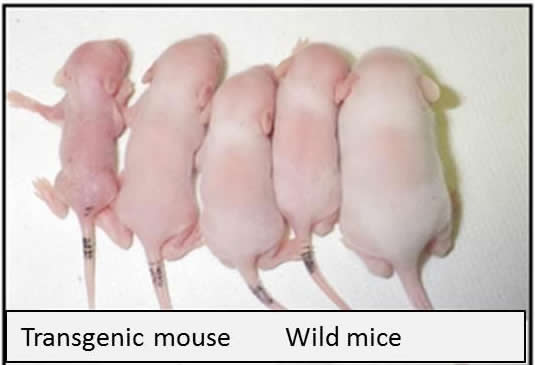
FURTHEST LEFT: a transgenic mouse and 4 RIGHT: wild mice.
Image Credit:
University of Tsukuba
|
|
| |
|



