|
|
Tuning the volume on gene expression
Research finds genes can be turned on and off, or finely adjusted as if controlled by a volume control knob.
Garth Ilsley PhD, research scientist in the Nicholas Luscombe laboratory at the Okinawa Institute of Science and Technology Graduate University (OIST), Onna, Japan, has developed a mathematic model to predict gene expression.
His experiments complement the pioneering work of Justin Crocker PhD, in the David Stern PhD, laboratory at Janelia Research Campus of the Howard Hughes Medical Institute, Ashburn, Virginia, USA. Published in Nature Genetics, the research has important implications in cellular and developmental biology including stem cell reprogramming and developments in regenerative medicine.
Transcription factors are proteins that bind to special regions of DNA called enhancers. They regulate gene expression. Some activate gene expression — turn genes on, while others repress gene expression — turn genes off. Some transcription factors also turn up performance or "turn up volume", while others turn performance down or "turn down volume" — like the knobs on a radio.
Scientists investigated how activation and repression work together to predict the potential level of gene expression. By turning the volume knob of the "gene radio", is it possible to regulate how loud or soft the result will be? In the case of genes, each transcription factor has its own volume knob — yet all act on the same speaker. The challenge was understanding how enhancers work together to produce one result.
The beauty of
Dr Ilsley's new mathematic model is that it does not require the number and position of each transcription factor as it is bound to an enhancer to work. Instead, his model predicts the final volume by only knowing how volume knobs work. The predictions were tested using artificial transcription factors in order to activate and repress gene expression at different signal strengths.
Working with fruit fly (Drosophila melanogaster) early embryos, the mathematic model reveals that turning on segmentation genes which determine body parts of the fruit fly from head to tail — is adjustable.
Experimental results matched their prediction.
With the mathematic model, activators (increased) and repressors (decreased) gene expression gradually and in a controllable and reproducible way. This result proved attaching a second volume knob to the radio automatically synchronised it to the existing volume knob.
Beyond gene expression, the mathematic model also predicted where the gene will be expressed — for the fly embryo — underneath (ventral) or on top (dorsal) of the fly body.
"It was our dream to bring model and experiment together," enthused Dr Ilsley. The results also showed that enhancers can acquire new activators and repressors with great flexibility.
"You can bring in foreign transcription factors and enhancers will still work. Enhancers we looked at are not brittle at all — this is evolutionarily important. It shows enhancer activity can be gradually adjusted and still work in spite of changing contexts.
"We are moving away from the on/off model of gene expression to explain how cell types become specific. Advances in quantitative biology at the single-cell level — together with mathematic models — give biologists tools to explore the intricacy of tuning gene expression — in order to predictably manipulate the cell."
Garth R. Ilsley PhD, laboratory at Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, Virginia, USA
Abstract
Genes are regulated by transcription factors that bind to regions of genomic DNA called enhancers. Considerable effort is focused on identifying transcription factor binding sites, with the goal of predicting gene expression from DNA sequence. Despite this effort, general, predictive models of enhancer function are currently lacking. Here we combine quantitative models of enhancer function with manipulations using engineered transcription factors to examine the extent to which enhancer function can be controlled in a quantitatively predictable manner. Our models, which incorporate few free parameters, can accurately predict the contributions of ectopic transcription factor inputs. These models allow the predictable 'tuning' of enhancers, providing a framework for the quantitative control of enhancers with engineered transcription factors.
Return to top of page
|
|
|
Feb 12, 2016 Fetal Timeline Maternal Timeline News News Archive
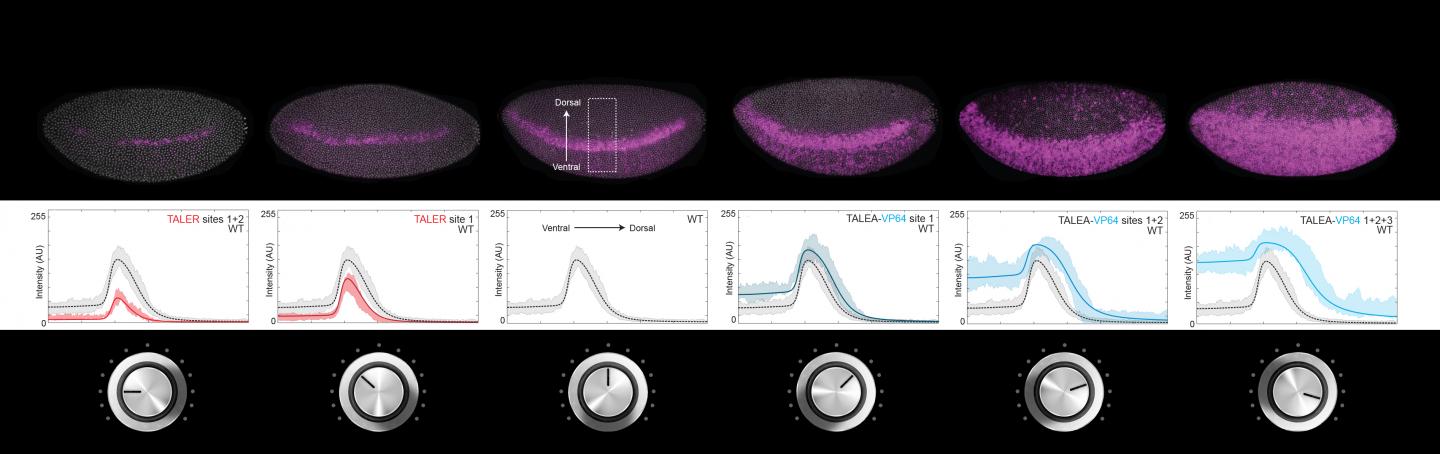
Different strengths of artificial transcription activators and repressors were introduced to fruit fly embryos and regulated how the gene "rhomboid" was expressed. Activators and repressors can
tune gene expression like volume knobs on a radio. Graphs predict the experimental results.
Image Credit: Modified from Nature Genetics
|
|
|
|



